This document assumes you know something about maximum likelihood estimation. It helps you get going in terms of doing MLE in R. All through this, we will use the "ordinary least squares" (OLS) model (a.k.a. "linear regression" or "classical least squares" (CLS)) as the simplest possible example. Here are the formulae for the OLS likelihood, and the notation that I use.
There are two powerful optimisers in R: optim() and nlminb(). This note only uses optim(). You should also explore nlminb().
You might find it convenient to snarf a tarfile of all the .R programs involved in this page.
You have to write an R function which computes out the likelihood function. As always in R, this can be done in several different ways.
One issue is that of restrictions upon parameters. When the search algorithm is running, it may stumble upon nonsensical values - such as a sigma below 0 - and you do need to think about this. One traditional way to deal with this is to "transform the parameter space". As an example, for all positive values of sigma, log(sigma) ranges from -infinity to +infinity. So it's safe to do an unconstrained search using log(sigma) as the free parameter.
Here is the OLS likelihood, written in a few ways.Confucius he said, when you write a likelihood function, do take the trouble of also writing it's gradient (the vector of first derivatives). You don't absolutely need it, but it's highly recommended. In my toy experiment, this seems to be merely a question of speed - using the analytical gradient makes the MLE go faster. But the OLS likelihood is unique and simple; it is globally quasiconcave and has a clear top. There could not be a simpler task for a maximisation routine. In more complex situations, numerical derivatives are known to give more unstable searches, while analytical derivatives give more reliable answers.
To use the other files on this page, you need to take my simulation setup file.
Now that I've written the OLS likelihood function in a few ways, it's natural to ask: Do they all give the same answer? And, which is the fastest?
I wrote a simple R program in order to learn these things. This gives the result:
True theta = 2 4 6
OLS theta = 2.004311 3.925572 6.188047
Kick the tyres --
lf1() lf1() in logs lf2() lf3()
A weird theta 1864.956 1864.956 1864.956 1864.956
True theta 1766.418 1766.418 1766.418 1766.418
OLS theta 1765.589 1765.589 1765.589 1765.589
Cost (ms) 0.450 0.550 1.250 1.000
Derivatives -- first let me do numerically --
Derivative in sigma -- 10.92756
Derivative in intercept -- -8.63967
Derivative in slope -- -11.82872
Analytical derivative in sigma -- 10.92705
Analytical derivative in beta -- -8.642051 -11.82950
This shows us that of the 4 ways of writing it, ols.lf1() is the fastest, and that there is a fair match between my claimed analytical gradient and numerical derivatives.
Using this foundation, I can jump to a self-contained and minimal R program which does the full job. It gives this result:
True theta = 2 4 6
OLS theta = 2.004311 3.925572 6.188047
Gradient-free (constrained optimisation) --
$par
[1] 2.000304 3.925571 6.188048
$value
[1] 1765.588
$counts
function gradient
18 18
$convergence
[1] 0
$message
[1] "CONVERGENCE: REL_REDUCTION_OF_F <= FACTR*EPSMCH"
Using the gradient (constrained optimisation) --
$par
[1] 2.000303 3.925571 6.188048
$value
[1] 1765.588
$counts
function gradient
18 18
$convergence
[1] 0
$message
[1] "CONVERGENCE: REL_REDUCTION_OF_F <= FACTR*EPSMCH"
You say you want a covariance matrix?
MLE results --
Coefficient Std. Err. t
Sigma 2.000303 0.08945629 22.36068
Intercept 3.925571 0.08792798 44.64530
X 6.188048 0.15377325 40.24138
Compare with the OLS results --
Estimate Std. Error t value Pr(>|t|)
(Intercept) 3.925572 0.08801602 44.60065 7.912115e-240
X[, 2] 6.188047 0.15392722 40.20112 6.703474e-211
The file minimal.R also generates this picture:
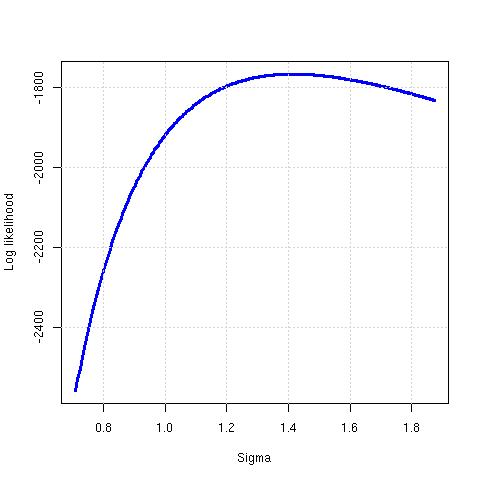
The R optim() function has many different paths to MLE. I wrote a simple R program in order to learn about these. This yields the result:
True theta = 2 4 6
OLS theta = 2.004311 3.925572 6.188047
Hit rate Cost
L-BFGS-B, analytical 100 25.1
BFGS, analytical 100 33.1
Nelder-Mead, trans. 100 59.2
Nelder-Mead 100 60.5
L-BFGS-B, numerical 100 61.2
BFGS, trans., numerical 100 68.5
BFGS, numerical 100 71.2
SANN 99 4615.5
SANN, trans. 96 4944.9
The algorithms compared above are:
I learned this with help from Douglas Bates and Spencer Graves, and by lurking on the r-help mailing list, thus learning from everyone who takes the trouble of writing there. Thanks!
Back up to my R knowledge base system
Ajay Shah
ajayshah at mayin dot org